Before doing first-level analysis, you must run brain extraction using BET, ensure your functional and structural data are in NIFTI format, and ensure that you have any necessary event timing files (e.g., stimulus onset, durations).
Launch Feat GUI:
Feat &The interactive interface of Feat from FSL:
Set Up the First-Level Analysis
Define the Data Input • Data Directory: Select the folder containing your functional data. • Data Type: Choose whether your design is “Time-series” (single run) or “Multiple sessions” (if you have more than one run).
Specify Preprocessing Options (if needed) • Motion Correction: Enable MCFLIRT if not already done. • Slice Timing Correction: Check this if your sequence requires correction (and if it wasn’t already applied). • Spatial Smoothing: Specify the smoothing kernel size (e.g., 5 mm FWHM). • High-Pass Filtering: Set the cutoff (e.g., 100 s) to remove low-frequency drifts.
Set Up the Model
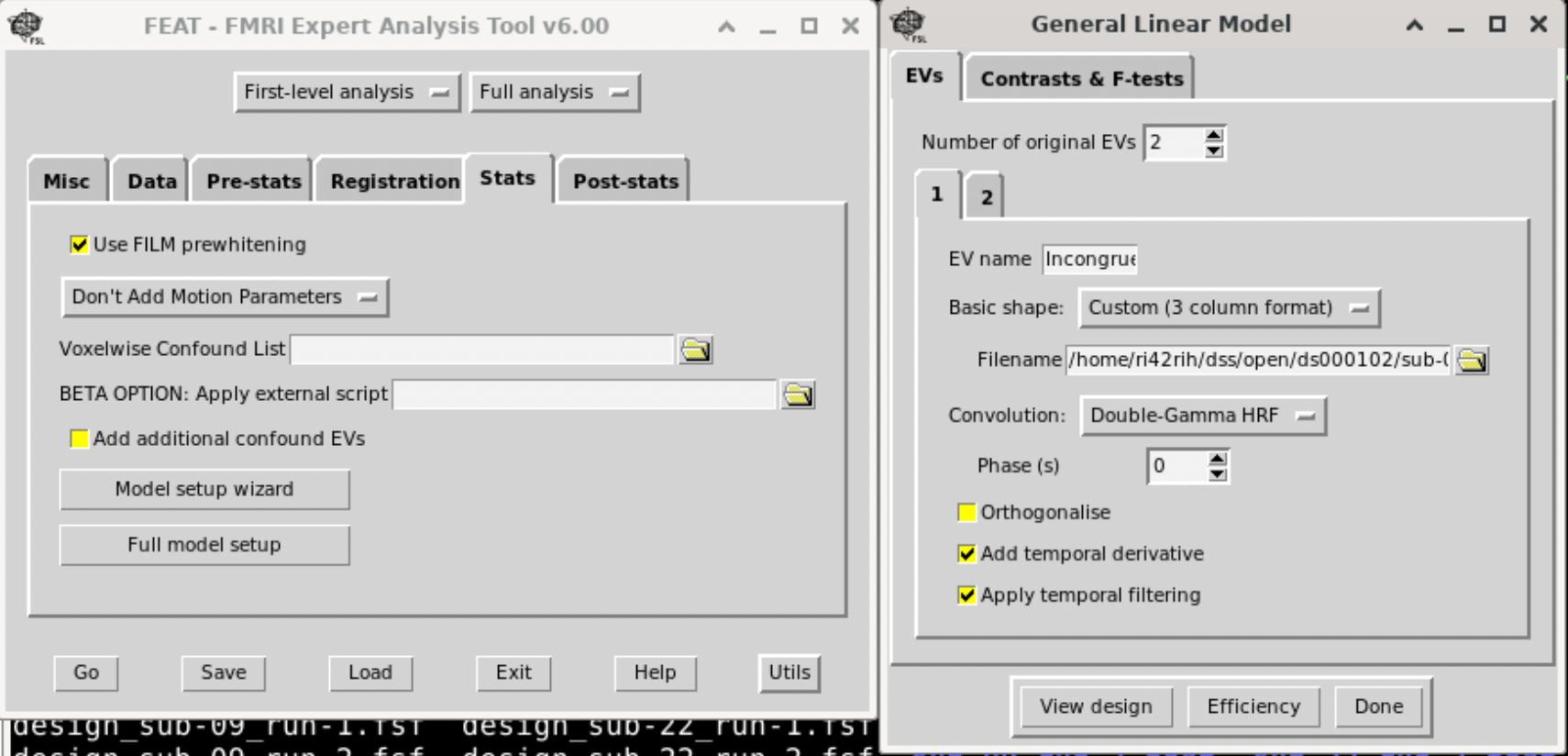
• Design Setup: • Click on “Stats” in the GUI. • EVs (Explanatory Variables): Define your regressors. For each experimental condition, you can either use a block or event-related model: • Click “Add EV” for each condition. • Provide the timing file or manually input the onsets/durations. Usually we provide a three-column text for this with the option ‘Custom (3 column format)‘. Three columns are: Onset, Duration, and amplitude (1).
• Temporal Derivatives (optional): Include them if you want to account for slight differences in response timing. • Contrasts: • Under the “Contrasts” tab, set up contrasts to test your hypotheses (e.g., condition A > baseline). • Add as many contrasts as needed to test different effects.
Registration
• Structural Registration: • Register your functional data to your high-resolution structural scan. • Use FLIRT for linear registration (and optionally FNIRT for nonlinear registration if needed). • Check the transformation matrices and overlay the images to verify alignment. • Standard Space Registration: If you plan to compare with group-level analyses, register to a standard template (e.g., MNI152 2mm).
Then click ‘Go’ to run the Analysis and save the template, which we need to adapt for other subjects.
Adapt template for all participants
The saved Feat configuration .fsf is actually a text file, which can be edited by any text editor. We open this saved file, and change the following fields to variables, which we will replace using a bash file:
- Output directory
- Total Volumes
- Functional image location
- T1 image location
- 3-column timing files The following code is an example of replaced template.
# Output directory
set fmri(outputdir) "OUTPUT_DIR"
# Total volumes
set fmri(npts) TOTAL_VOLUMES
# 4D AVW data or FEAT directory (1)
set feat_files(1) "FUNC_IMAGE"
# Subject's structural image for analysis 1
set highres_files(1) "T1_IMAGE"
# Custom EV file (EV 1)
set fmri(custom1) "COND1_FILE"
# Custom EV file (EV 2)
set fmri(custom2) "COND2_FILE"
If you have multiple RUNs, you should also make them replaceable variables.
Next, use a bash file to replace them to run Feat for individual subjects.
#!/bin/bash
SCRIPT_DIR=$(cd "$(dirname "$0")" && pwd)
BIDS_DIR=$(cd "$SCRIPT_DIR/../.." && pwd)
DERIVATIVES_DIR="${BIDS_DIR}/derivatives" # Output directory for first-level results
FSF_TEMPLATE="${BIDS_DIR}/code/fsl/template_level.fsf"
# Ensure the derivatives directory exists
mkdir -p "${DERIVATIVES_DIR}"
# Number of parallel jobs to run
NJOBS=8
# Track jobs to prevent overload
job_count=0
# Loop over all subject folders
for subj in ${BIDS_DIR}/sub-*; do
subj_id=$(basename "${subj}") # e.g., sub-01
func_dir="${subj}/func"
# Locate the T1 brain image (common to all runs)
t1_image=$(ls "${subj}/anat/${subj_id}_T1w_brain.nii.gz" 2>/dev/null | head -n 1)
# Loop over runs (e.g., run-1 and run-2)
# depending how many runs you have
for run in 1 2; do
# Locate functional image for this run
func_image=$(ls "${func_dir}/${subj_id}_task-flanker_run-${run}_bold.nii.gz" 2>/dev/null | head -n 1)
echo "${func_dir}/${subj_id}_task-*_run-${run}_bold.nii.gz"
# Locate the event files for congruent and incongruent conditions for this run
cond2_file=$(ls "${func_dir}/congruent_run${run}.txt" 2>/dev/null | head -n 1)
cond1_file=$(ls "${func_dir}/incongruent_run${run}.txt" 2>/dev/null | head -n 1)
# Skip if no functional image is found for this run
if [ ! -f "$func_image" ]; then
echo "No functional image found for ${subj_id} run-${run}, skipping..."
continue
fi
# Get the number of volumes (4th dimension) from the functional image
total_vols=$(fslval "$func_image" dim4)
# Define run-specific FSF file and FEAT output directory
fsf_file="${DERIVATIVES_DIR}/fsl/design_${subj_id}_run-${run}.fsf"
output_dir="${DERIVATIVES_DIR}/fsl/${subj_id}_run-${run}.feat"
echo "output_dir: ${output_dir}"
echo "fsf_file: ${fsf_file}"
# Skip if FEAT analysis has already been run for this run
if [ -d "${output_dir}" ]; then
echo "Skipping ${subj_id} run-${run}, FEAT already run."
continue
fi
# Create a subject- and run-specific FSF file with updated paths and parameters.
# Note: The FSF template should include placeholders such as SUBJ-ID, RUN_NUMBER,
# OUTPUT_DIR, FUNC_IMAGE, T1_IMAGE, COND1_FILE, COND2_FILE, and TOTAL_VOLUMES.
sed -e "s/SUBJ-ID/${subj_id}/g" \
-e "s/RUN_NUMBER/run-${run}/g" \
-e "s|OUTPUT_DIR|${output_dir}|g" \
-e "s|FUNC_IMAGE|${func_image}|g" \
-e "s|T1_IMAGE|${t1_image}|g" \
-e "s|COND1_FILE|${cond1_file}|g" \
-e "s|COND2_FILE|${cond2_file}|g" \
-e "s/TOTAL_VOLUMES/${total_vols}/g" \
"${FSF_TEMPLATE}" > "${fsf_file}"
# Run FEAT for this subject and run in the background
echo "Running FEAT for ${subj_id} run-${run} with ${total_vols} volumes..."
feat "${fsf_file}" &
# Track the number of background jobs and wait if the limit is reached
job_count=$((job_count + 1))
if [ $job_count -eq $NJOBS ]; then
wait
job_count=0
fi
done
done
# Wait for any remaining background jobs to complete
wait
echo "First-level FEAT analysis completed for all subjects and runs."