Analysis Overview
This tutorial implements the standard 3-level fMRI analysis hierarchy:
- Run-level (1st level): GLM for each individual run → COPEs and VARCOPEs
- Subject-level (fixed-effects): Combine runs within subjects → Fixed-effects COPEs
- Group-level (2nd level/random-effects): Model population using subject-level COPEs → Group statistics
Key principle: We propagate effect sizes (COPEs), not standardized statistics (z-maps), up the hierarchy. This preserves effect magnitudes and allows proper random-effects modeling across subjects.
0 Prerequisites
| Requirement | Why you need it |
|---|---|
| Python ≥ 3.9 with nilearn 0.12+, numpy, pandas, scipy, nibabel, joblib | Core libraries |
| fMRIPrep-pre-processed data (BIDS derivatives) | Gives you: *_desc-preproc_bold.nii.gz, *_desc-brain_mask.nii.gz, *_events.tsv, *_confounds_timeseries.tsv |
| An MNI brain mask (e.g. MNI152_T1_2mm_brain_mask.nii.gz) | For group-level masking |
Install everything (one-off):
pip install nilearn nibabel scipy pandas joblib
1 Per-run first-level GLM
import os, glob, pandas as pd
from nilearn.glm.first_level import FirstLevelModel
proj_dir = '/path/to/dataset'
results_dir = '/path/to/level1_nilearn'
subjects = sorted([d.split('-')[1] for d in glob.glob(f'{proj_dir}/sub-*')])
runs = ['1', '2']
contrasts = {
'Congruent' : 'congruent',
'Incongruent' : 'incongruent',
'Incongr>Cong': 'incongruent - congruent'
}
motion_cols = ['rot_x','rot_y','rot_z','trans_x','trans_y','trans_z']
for sub in subjects:
for run in runs:
# ----- file paths ---------------------------------------------------
func = f'{proj_dir}/derivatives/fmriprep/sub-{sub}/func/' \
f'sub-{sub}_task-flanker_run-{run}_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz'
mask = func.replace('preproc_bold', 'brain_mask')
events = f'{proj_dir}/sub-{sub}/func/' \
f'sub-{sub}_task-flanker_run-{run}_events.tsv'
confounds_tsv = f'{proj_dir}/derivatives/fmriprep/sub-{sub}/func/' \
f'sub-{sub}_task-flanker_run-{run}_desc-confounds_timeseries.tsv'
events_df = pd.read_csv(events, sep='\t')
confounds_df = pd.read_csv(confounds_tsv, sep='\t')[motion_cols]
# ----- model --------------------------------------------------------
model = FirstLevelModel(
t_r=2.0, smoothing_fwhm=6.0, mask_img=mask,
drift_model='cosine', high_pass=1/128, noise_model='ar1'
).fit(func, events=events_df, confounds=confounds_df)
out_dir = f'{results_dir}/sub-{sub}/run-{run}'
os.makedirs(out_dir, exist_ok=True)
for name, expr in contrasts.items():
model.compute_contrast(expr, output_type='effect_size' ).to_filename(f'{out_dir}/{name}_cope.nii.gz')
model.compute_contrast(expr, output_type='effect_variance').to_filename(f'{out_dir}/{name}_varcope.nii.gz')
model.compute_contrast(expr, output_type='z_score' ).to_filename(f'{out_dir}/{name}_zmap.nii.gz')What happened?
- Design matrix construction: Events are convolved with HRF, combined with motion regressors
- Model fitting: One call to fit() performs design-matrix creation, HRF-convolution, and GLS estimation
- Contrast computation: For each contrast, we compute:
- COPE (effect_size): The contrast estimate (c’β) - this is what we need for group analysis
- VARCOPE (effect_variance): The variance of the contrast estimate - needed for fixed-effects
- Z-map (z_score): Standardized statistic for this run only (COPE/sqrt(VARCOPE))
2 Within-subject fixed-effects (combine runs)
from nilearn.glm import compute_fixed_effects
subject_copes = {} # {sub: {contrast: fixed-effects cope img}}
for sub in subjects:
subject_copes[sub] = {}
for c in contrasts:
copes = [f'{results_dir}/sub-{sub}/run-{r}/{c}_cope.nii.gz' for r in runs]
varcps = [f'{results_dir}/sub-{sub}/run-{r}/{c}_varcope.nii.gz' for r in runs]
fx_cope, fx_var, _, fx_z = compute_fixed_effects(
copes, varcps, precision_weighted=True, return_z_score=True
)
# Save the fixed-effects contrast estimate (effect size) for group analysis
cope_fout = f'{results_dir}/sub-{sub}/{c}_fixed_cope.nii.gz'
fx_cope.to_filename(cope_fout)
subject_copes[sub][c] = cope_fout
# Also save z-map and variance for completeness/diagnostic purposes
fx_z.to_filename(f'{results_dir}/sub-{sub}/{c}_fixed_zmap.nii.gz')
fx_var.to_filename(f'{results_dir}/sub-{sub}/{c}_fixed_varcope.nii.gz')What happened?
- compute_fixed_effects does an inverse-variance weighted average of runs
- Key point: We save the fixed-effects contrast estimates (COPEs) for group analysis, not z-maps
- Z-maps represent standardized effects specific to each subject’s noise level and are not appropriate for group-level random-effects analysis
3 Group (second-level) random-effects
from nilearn.glm.second_level import SecondLevelModel
import pandas as pd
group_dir = f'{results_dir}/group'
os.makedirs(group_dir, exist_ok=True)
for c in contrasts:
# Use contrast estimates (effect sizes), NOT z-maps, for group analysis
inputs = [subject_copes[sub][c] for sub in subjects]
design = pd.DataFrame({'intercept': [1]*len(inputs)})
model = SecondLevelModel(smoothing_fwhm=6.0, mask_img='MNI152_T1_2mm_brain_mask.nii.gz')
model = model.fit(inputs, design)
# Generate group-level statistics
cope = model.compute_contrast('intercept', output_type='effect_size')
zmap = model.compute_contrast('intercept', output_type='z_score')
tmap = model.compute_contrast('intercept', output_type='stat')
# Save outputs
cope.to_filename(f'{group_dir}/{c}_group_cope.nii.gz')
zmap_fn= f'{group_dir}/{c}_group_zmap.nii.gz'
zmap.to_filename(zmap_fn)
tmap.to_filename(f'{group_dir}/{c}_group_tmap.nii.gz')Why use contrast estimates (COPEs) instead of z-maps?
- Random-effects analysis requires effect sizes that can be averaged across subjects
- Z-maps are standardized by each subject’s individual error variance - they don’t preserve the actual magnitude of effects
- COPEs represent the actual c’β (contrast times beta) values that can be meaningfully compared across subjects
4 Cluster-level inference + table
from nilearn.glm import cluster_level_inference
from nilearn.reporting import get_clusters_table
from scipy.stats import norm
z_thresh = norm.isf(0.001) # voxel p<0.001
zmap_file = f'{group_dir}/Incongr>Cong_group_zmap.nii.gz'
# A. cluster FWE p<0.05
clusters_img = cluster_level_inference(
zmap_file, mask_img='MNI152_T1_2mm_brain_mask.nii.gz',
threshold=z_thresh, alpha=0.05
)
clusters_img.to_filename(f'{group_dir}/Incongr>Cong_clusterFWE_zmap.nii.gz')
# B. cluster table (≥20 vox @ voxel p<0.001)
table = get_clusters_table(zmap_file, stat_threshold=z_thresh, cluster_threshold=20)
print(table.head())5 Visualisation
from nilearn import plotting
from nilearn.image import index_img
from nilearn.glm.thresholding import threshold_stats_img
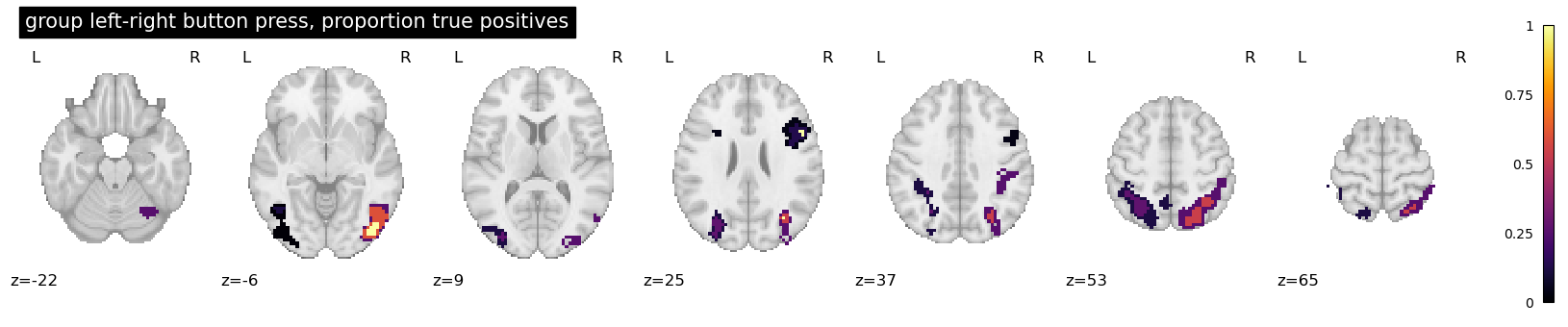
plotting.plot_stat_map(
clusters_img,
threshold=0.0, display_mode="z", vmax=1,
cmap="inferno",
title="group left-right button press, proportion true positives",
)
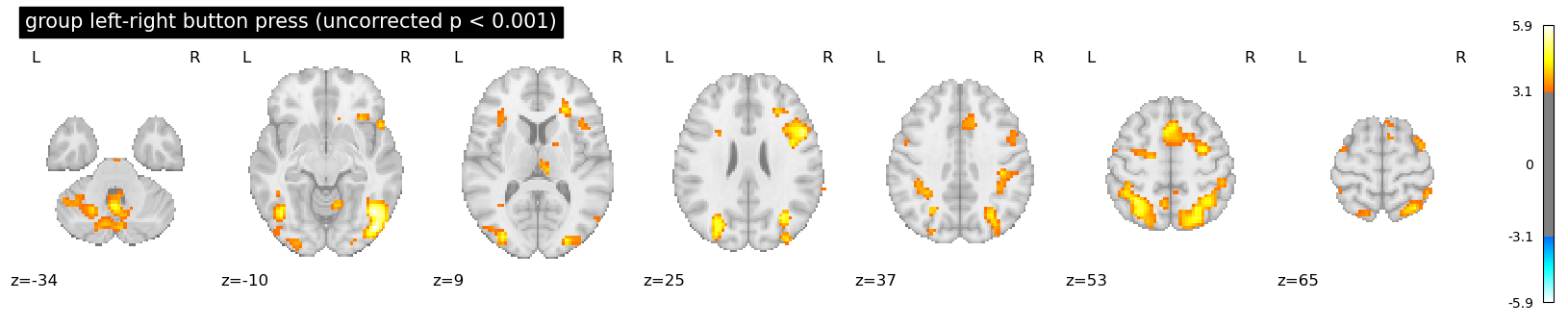
plotting.plot_stat_map(
zmap_file,
threshold=z_thresh_uncorr,
display_mode="z",
title="group left-right button press (uncorrected p < 0.001)",
)
6 Parallelising (optional)
from joblib import Parallel, delayed
Parallel(n_jobs=6)(
delayed(process_run)(sub, run, ...) # wrap the first-level code in a function
for sub in subjects for run in runs
)
7 Summary
| Step | Nilearn function(s) | What it replaces in FSL/SPM/AFNI |
|---|---|---|
| 1 | FirstLevelModel | FEAT (FSL), 3dDeconvolve (AFNI), SPM first-level |
| 2 | compute_fixed_effects | FEAT “higher-level” fixed-effects |
| 3 | SecondLevelModel | FLAME, 3dttest++, SPM second-level |
| 4 | cluster_level_inference, get_clusters_table | cluster, randomise, 3dClustSim |
| 5 | plot_glass_brain, view_img_on_surf | FSLeyes, AFNI viewer, SPM GUI |
Further reading
-
Nilearn docs: https://nilearn.github.io
-
GLM tutorial: https://nilearn.github.io/dev/auto_examples/05_glm_first_level
-
Cluster inference: Rosenblatt et al., 2018, NeuroImage